Problems¶
This section covers Problem classes in more detail.
Class Summary¶
Figure 1: The `Problem` abstract-base class This class diagram shows the detail for Problem, which is an abstract base class (ABC). It has three abstract methods that must be over-ridden by subclasses. evaluate() takes a phenome from an individual and compute a fitness from that. worse_than() and equivalent() compare fitnesses from two different individuals and, as the name suggests, respectively returns the worst of the two or the equivalent within the Problem context.¶
As shown in Fig. 1, the Problem`abstract-base class has three abstract methods. `evaluate() takes a phenome that was decode()d from an Individual’s genome, and returns a value denoting the quality, or fitness, of that individual. Problems are also used to compare the fitnesses between Individuals. worse_than() returns true if the first individual is less fit than the second. Similarly, equivalent() is used to determine if two given fitnesses are effectively the same.
Class API¶

Defines the abstract-base classes Problem, ScalarProblem, and FunctionProblem.
-
class
leap_ec.problem.ConstantProblem(maximize=False, c=1.0)¶ A flat landscape, where all phenotypes have the same fitness.
This is sometimes useful for sanity checks or as a control in certain kinds of research.
\[f(\vec{x}) = c\]- Parameters
c (float) – the fitness value to return for any input.
from leap_ec.problem import ConstantProblem from leap_ec.real_rep.problems import plot_2d_problem bounds = ConstantProblem.bounds plot_2d_problem(ConstantProblem(), xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)

-
bounds= (-1.0, 1.0)¶
-
evaluate(phenome, *args, **kwargs)¶ Return a contant value for any input phenome:
>>> phenome = [0.5, 0.8, 1.5] >>> ConstantProblem().evaluate(phenome) 1.0
>>> ConstantProblem(c=500.0).evaluate('foo bar') 500.0
- Parameters
phenome – real-valued vector to be evaluated
- Returns
1.0, or the constant defined in the constructor
-
class
leap_ec.problem.FunctionProblem(fitness_function, maximize)¶ -
evaluate(phenome, *args, **kwargs)¶ Decode and evaluate the given individual based on its genome.
Practitioners must over-ride this member function.
Note that by default the individual comparison operators assume a maximization problem; if this is a minimization problem, then just negate the value when returning the fitness.
- Parameters
phenome –
- Returns
fitness
-
-
class
leap_ec.problem.Problem¶ Abstract Base Class used to define problem definitions.
A Problem is in charge of two major parts of an EA’s behavior:
Fitness evaluation (the evaluate() method)
Fitness comparision (the worse_than() and equivalent() methods)
-
abstract
equivalent(first_fitness, second_fitness)¶
-
abstract
evaluate(phenome, *args, **kwargs)¶ Decode and evaluate the given individual based on its genome.
Practitioners must over-ride this member function.
Note that by default the individual comparison operators assume a maximization problem; if this is a minimization problem, then just negate the value when returning the fitness.
- Parameters
phenome –
- Returns
fitness
-
abstract
worse_than(first_fitness, second_fitness)¶
-
class
leap_ec.problem.ScalarProblem(maximize)¶ -
equivalent(first_fitness, second_fitness)¶ Used in Individual.__eq__().
By default returns first.fitness== second.fitness. Please over-ride if this does not hold for your problem.
- Returns
true if the first individual is equal to the second
-
worse_than(first_fitness, second_fitness)¶ Used in Individual.__lt__().
By default returns first_fitness < second_fitness if a maximization problem, else first_fitness > second_fitness if a minimization problem. Please over-ride if this does not hold for your problem.
- Returns
true if the first individual is less fit than the second
-
Binary Problems API¶

A set of standard EA problems that rely on a binary-representation
-
class
leap_ec.binary_rep.problems.ImageProblem(path, maximize=True, size=100, 100)¶ A variation on max_ones that uses an external image file to define a binary target pattern.
-
evaluate(phenome)¶ Decode and evaluate the given individual based on its genome.
Practitioners must over-ride this member function.
Note that by default the individual comparison operators assume a maximization problem; if this is a minimization problem, then just negate the value when returning the fitness.
- Parameters
phenome –
- Returns
fitness
-
-
class
leap_ec.binary_rep.problems.MaxOnes(maximize=True)¶ Implementation of MAX ONES problem where the individuals are represented by a bit vector
We don’t need an encoder since the raw genome is already in the phenotypic space.
-
evaluate(phenome)¶ >>> from leap_ec.individual import Individual >>> from leap_ec.decoder import IdentityDecoder >>> p = MaxOnes() >>> ind = Individual([0, 0, 1, 1, 0, 1, 0, 1, 1], ... decoder=IdentityDecoder(), ... problem=p) >>> p.evaluate(ind.decode()) 5
-
Real-value Problems API¶

This module contains a variety of classic real-valued optimization problems that frequently occur in research benchmarks.
It also contains helpers for translating, rotating, and visualizing them.
-
class
leap_ec.real_rep.problems.AckleyProblem(a=20, b=0.2, c=6.283185307179586, maximize=False)¶ - \[f(\mathbf{x}) = -a \exp \left( -b \sqrt \frac{1}{d} \sum_{i=1}^d x_i^2 \right) - \exp \left( \frac{1}{d} \sum_{i=1}^d \cos(cx_i) \right) + a + \exp(1)\]
- Parameters
a (float) – depth parameter for the bowl-shaped macrostructure
b (float) – exponential scale parameter for the bowl
c (float) – wavenumber (frequency) of the cosine pattern of local optima
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import AckleyProblem, plot_2d_problem import math problem = AckleyProblem(a=20, b=0.2, c=2*math.pi) bounds = AckleyProblem.bounds # Contains traditional bounds plot_2d_problem(problem, xlim=bounds, ylim=bounds, granularity=0.25)
(Source code, png, hires.png, pdf)
-
bounds= [-32.768, 32.768]¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
class
leap_ec.real_rep.problems.CosineFamilyProblem(alpha, global_optima_counts, local_optima_counts, maximize=False)¶ A configurable multi-modal function based on combinations of cosines, taken from the problem generators proposed in
- Jani2008
“A Generator for Multimodal Test Functions with Multiple Global Optima,” Jani Rönkkönen et al., Asia-Pacific Conference on Simulated Evolution and Learning. Springer, Berlin, Heidelberg, 2008.
\[f_{\cos}(\mathbf{x}) = \frac{\sum_{i=1}^n -\cos((G_i - 1)2 \pi x_i) - \alpha \cdot \cos((G_i - 1)2 \pi L-i x_y)}{2n}\]where \(G_i\) and \(L_i\) are parameters that indicate the number of global and local optima, respectively, in the ith dimension.
- Parameters
alpha (float) – parameter that controls the depth of the local optima.
global_optima_counts ([int]) – list of integers indicating the number of global optima for each dimension.
local_optima_counts ([int]) – list of integers indicated the number of local optima for each dimension.
maximize – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import CosineFamilyProblem, plot_2d_problem problem = CosineFamilyProblem(alpha=1.0, global_optima_counts=[2, 2], local_optima_counts=[2, 2]) bounds = CosineFamilyProblem.bounds # Contains traditional bounds plot_2d_problem(problem, xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)
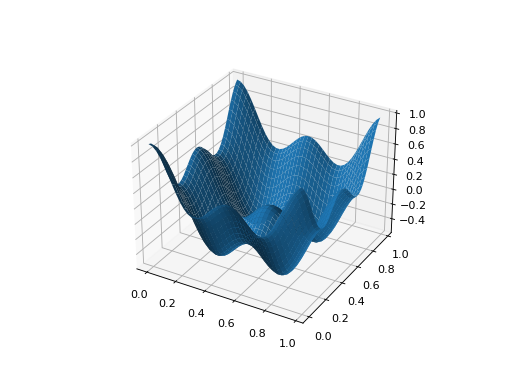
The number of optima can be varied independently by each dimension:
from leap_ec.real_rep.problems import CosineFamilyProblem, plot_2d_problem problem = CosineFamilyProblem(alpha=3.0, global_optima_counts=[4, 2], local_optima_counts=[2, 2]) bounds = CosineFamilyProblem.bounds # Contains traditional bounds plot_2d_problem(problem, xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)
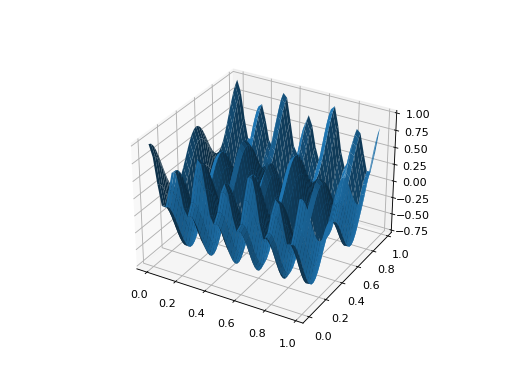
-
bounds= (0, 1)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
class
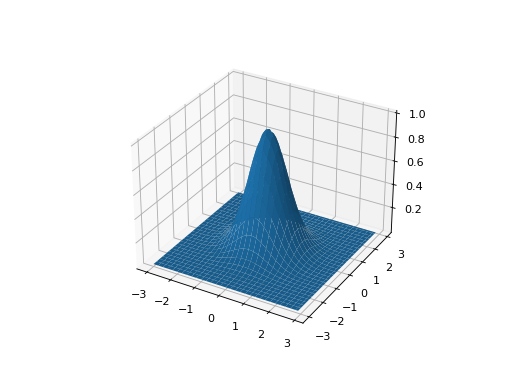
leap_ec.real_rep.problems.GaussianProblem(width=1, height=1, maximize=True)¶ A multidimensional, isotropic Gaussian function, defined by
\[A\exp\left( - \sum_i^n \left(\frac{x_i}{w}\right)^2 \right)\]- Parameters
width (float) – the width parameter \(w\)
height (float) – the height parameter \(A\)
from leap_ec.real_rep.problems import GaussianProblem, plot_2d_problem bounds = GaussianProblem.bounds # Some typical bounds problem = GaussianProblem(width=1, height=1) plot_2d_problem(problem, xlim=bounds, ylim=bounds, granularity=0.1)
(Source code, png, hires.png, pdf)
-
bounds= (-3, 3)¶
-
evaluate(phenome)¶ Decode and evaluate the given individual based on its genome.
Practitioners must over-ride this member function.
Note that by default the individual comparison operators assume a maximization problem; if this is a minimization problem, then just negate the value when returning the fitness.
- Parameters
phenome –
- Returns
fitness
-
class
leap_ec.real_rep.problems.GriewankProblem(maximize=False)¶ The classic Griewank problem. Like the
RastriginProblemfunction, the Griewank has a quadratic global structure with many local optima that are distributed in a regular pattern.\[f(\mathbf{x}) = \sum_{i=1}^d \frac{x_i^2}{4000} - \prod_{i=1}^d \cos\left(\frac{x_i}{\sqrt{i}}\right) + 1\]- Parameters
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import GriewankProblem, plot_2d_problem bounds = GriewankProblem.bounds # Contains traditional bounds plot_2d_problem(GriewankProblem(), xlim=bounds, ylim=bounds, granularity=10)
(Source code, png, hires.png, pdf)

from leap_ec.real_rep.problems import GriewankProblem, plot_2d_problem bounds = [-50, 50] plot_2d_problem(GriewankProblem(), xlim=bounds, ylim=bounds, granularity=1)
(Source code, png, hires.png, pdf)

-
bounds= [-600, 600]¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
class
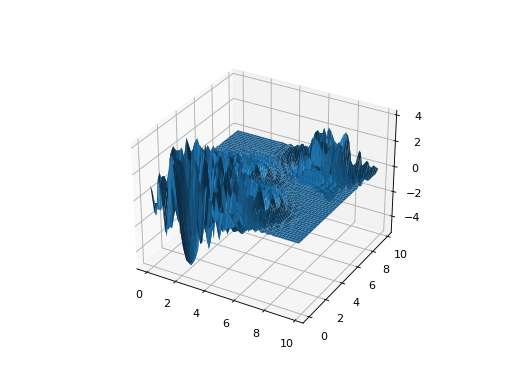
leap_ec.real_rep.problems.LangermannProblem(m=5, c=1, 2, 5, 2, 3, a=3, 5, 5, 2, 2, 1, 1, 4, 7, 9, maximize=False)¶ A popular multi-modal test function built by summing together \(m\) terms.
\[f(\mathbf{x}) = -\sum_{i=1}^m c_i \exp\left( -\frac{1}{\pi} \sum_{j=1}^d(x_j - A_{ij})^2\right) \cos\left(\pi\sum_{j=1}^d(x_j - A_{ij})^2\right)\]Langermann’s function is parameterized by a vector \(c_i\) of length \(m\) and a matrix \(A_{ij}\) of dimension \(m \times d\). This class uses the traditional parameterization as the default, with \(m=5\) and
\[\begin{split}c = (1, 2, 5, 2, 3) \\ A = \left[ \begin{array}{ll} 3 & 5\\ 5 & 2\\ 2 & 1\\ 1 & 4\\ 7 & 9 \end{array} \right].\end{split}\]- Parameters
m (int) – total number of terms in the function’s sum
c ([float]) – amplitude coefficients for each term
a ([[float]]) – offsets points for each term
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import LangermannProblem, plot_2d_problem bounds = LangermannProblem.bounds # Contains traditional bounds plot_2d_problem(LangermannProblem(), xlim=bounds, ylim=bounds, granularity=0.2)
(Source code, png, hires.png, pdf)
-
bounds= [0, 10]¶
-
default_a= ((3, 5), (5, 2), (2, 1), (1, 4), (7, 9))¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
class
leap_ec.real_rep.problems.LunacekProblem(N, d=1.0, mu_1=2.5, mu_2=None, s=None, maximize=False)¶ Lunacek’s function is also know as the “double Rastrigin” or “bi-Rastrigin” problem, because it overlays a
RastriginProblem-style cosine function across a pair of spheroid functions.This function was designed to model the double-funnel macrostructure that occurs in some difficult cases of the Lennard-Jones function (a famous function from molecular dynamics).
\[f(\mathbf{x}) = \min \left( \left\{ \sum_{i=1}^N(x_i - \mu_1)^2 \right\}, \left\{ d \cdot N + s \cdot \sum_{i=1}^N(x_i - \mu_2)^2\right\} \right) + 10\sum_{i=1}^N(1 - \cos(2\pi(x_i-\mu_i))),\]where \(N\) is the dimensionality of the solution vector, and the second sphere center parameter \(\mu_2\) is typically given by
\[\mu_2 = -\sqrt{\frac{\mu_1^2 - d}{s}}\]and \(s\) is by default a function on \(N\):
\[s = 1 - \frac{1}{2\sqrt{N + 20} - 8.2}\]These respective defaults are used for \(\mu_2\) and \(s\) whenever mu_2 and s are set to None.
Because of these complicated defaults, this class requires that you explicitly set the dimensionality of \(N\) of the expected input solutions. A warning will be thrown if an input solution is encountered that doesn’t match the expected dimensionality.
- Parameters
N (int) – dimensionality of the anticipated input solutions
d (float) – base fitness value of the second spheroid
mu_1 (float) – offset of the first spheroid
mu_2 (float) – offset of the second spheroid (if None, this will be calculated automatically)
s (float) – scale parameter for the second spheroid (if None, this will be calculated automatically)
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import LunacekProblem, plot_2d_problem bounds = LunacekProblem.bounds # Contains traditional bounds plot_2d_problem(LunacekProblem(N=2), xlim=bounds, ylim=bounds, granularity=0.1)
(Source code, png, hires.png, pdf)

-
bounds= (-5, 5)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
class
leap_ec.real_rep.problems.MatrixTransformedProblem(problem, matrix, maximize=None)¶ Apply a linear transformation to a fitness function.
- Parameters
matrix – an nxn matrix, where n is the genome length.
- Returns
a function that first applies -matrix to the input, then applies fun to the transformed input.
For example, here we manually construct a 2x2 rotation matrix and apply it to the
leap.RosenbrockProblemfunction:from matplotlib import pyplot as plt from leap_ec.real_rep.problems import RosenbrockProblem, MatrixTransformedProblem, plot_2d_problem original_problem = RosenbrockProblem() theta = np.pi/2 matrix = [[np.cos(theta), -np.sin(theta)], [np.sin(theta), np.cos(theta)]] transformed_problem = MatrixTransformedProblem(original_problem, matrix) fig = plt.figure(figsize=(12, 8)) plt.subplot(221, projection='3d') bounds = RosenbrockProblem.bounds # Contains traditional bounds plot_2d_problem(original_problem, xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(222, projection='3d') plot_2d_problem(transformed_problem, xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(223) plot_2d_problem(original_problem, kind='contour', xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(224) plot_2d_problem(transformed_problem, kind='contour', xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025)
(Source code, png, hires.png, pdf)

-
evaluate(phenome)¶ Evaluated the fitness of a point on the transformed fitness landscape.
For example, consider a sphere function whose global optimum is situated at (0, 1):
>>> s = TranslatedProblem(SpheroidProblem(), offset=[0, 1]) >>> round(s.evaluate([0, 1]), 5) 0
Now let’s take a rotation matrix that transforms the space by pi/2 radians:
>>> import numpy as np >>> theta = np.pi/2 >>> matrix = [[np.cos(theta), -np.sin(theta)], [np.sin(theta), np.cos(theta)]] >>> r = MatrixTransformedProblem(s, matrix)
The rotation has moved the new global optimum to (1, 0)
>>> round(r.evaluate([1, 0]), 5) 0.0
The point (0, 1) lies at a distance of sqrt(2) from the new optimum, and has a fitness of 2:
>>> round(r.evaluate([0, 1]), 5) 2.0
-
classmethod
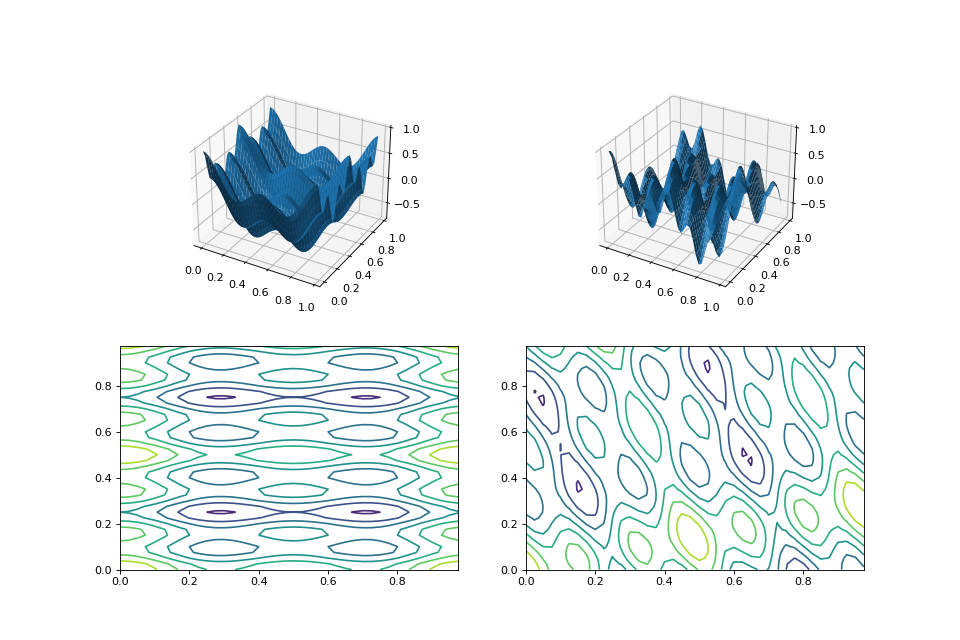
random_orthonormal(problem, dimensions, maximize=None)¶ Create a
MatrixTransformedProblemthat performs a random rotation and/or inversion of the function.We accomplish this by generating a random orthonormal basis for R^n and plugging the resulting matrix into
MatrixTransformedProblem.The classic algorithm we use here is based on the Gramm-Schmidt process: we first generate a set of random vectors, and then convert them into an orthonormal basis. This approach is described in Hansen and Ostermeier’s original CMA-ES paper:
“Completely derandomized self-adaptation in evolution strategies.” Evolutionary Computation 9.2 (2001): 159-195.
- Parameters
problem – the original
ScalarProblemto apply the transform to.dimensions (int) – the number of elements each vector should have.
maximize (bool) – whether to maximize or minimize the resulting fitness function. Defaults to whatever setting the underlying problem uses.
from matplotlib import pyplot as plt from leap_ec.real_rep.problems import CosineFamilyProblem, MatrixTransformedProblem, plot_2d_problem original_problem = CosineFamilyProblem(alpha=1.0, global_optima_counts=[2, 3], local_optima_counts=[2, 3]) transformed_problem = MatrixTransformedProblem.random_orthonormal(original_problem, 2) fig = plt.figure(figsize=(12, 8)) plt.subplot(221, projection='3d') bounds = original_problem.bounds plot_2d_problem(original_problem, xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(222, projection='3d') plot_2d_problem(transformed_problem, xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(223) plot_2d_problem(original_problem, kind='contour', xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(224) plot_2d_problem(transformed_problem, kind='contour', xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025)
(Source code, png, hires.png, pdf)
-
class
leap_ec.real_rep.problems.NoisyQuarticProblem(maximize=False)¶ The classic ‘quadratic quartic’ function with Gaussian noise:
\[f(\mathbf{x}) = \sum_{i=1}^{n} i x_i^4 + \texttt{gauss}(0, 1)\]- Parameters
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import NoisyQuarticProblem, plot_2d_problem bounds = NoisyQuarticProblem.bounds # Contains traditional bounds plot_2d_problem(NoisyQuarticProblem(), xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)

-
bounds= (-1.28, 1.28)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued list phenome (the output varies, since the function has noise):
>>> phenome = [3.5, -3.8, 5.0] >>> r = NoisyQuarticProblem().evaluate(phenome) >>> print(f'Result: {r}') Result: ...
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness
-
worse_than(first_fitness, second_fitness)¶ We minimize by default:
>>> s = NoisyQuarticProblem() >>> s.worse_than(100, 10) True
>>> s = NoisyQuarticProblem(maximize=True) >>> s.worse_than(100, 10) False
-
class
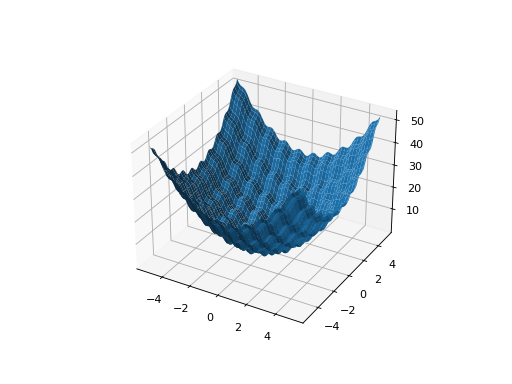
leap_ec.real_rep.problems.RastriginProblem(a=1.0, maximize=False)¶ The classic Rastrigin problem. The Rastrigin provides a real-valued fitness landscape with a quadratic global structure (like the
SpheroidProblem), plus a sinusoidal local structure with many local optima.\[f(\vec{x}) = An + \sum_{i=1}^n x_i^2 - A\cos(2\pi x_i)\]- Parameters
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import RastriginProblem, plot_2d_problem bounds = RastriginProblem.bounds # Contains traditional bounds plot_2d_problem(RastriginProblem(), xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)
-
bounds= (-5.12, 5.12)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued list phenome:
>>> phenome = [1.0/12, 0] >>> RastriginProblem().evaluate(phenome) # +doctest: ELLIPSIS 0.1409190406...
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness
-
worse_than(first_fitness, second_fitness)¶ We minimize by default:
>>> s = NoisyQuarticProblem() >>> s.worse_than(100, 10) True
>>> s = NoisyQuarticProblem(maximize=True) >>> s.worse_than(100, 10) False
-
class
leap_ec.real_rep.problems.RosenbrockProblem(maximize=False)¶ The classic RosenbrockProblem problem, a.k.a. the “banana” or “valley” function.
\[f(\mathbf{x}) = \sum_{i=1}^{d-1} \left[ 100 (x_{i + 1} - x_i^2)^2 + (x_i - 1)^2\right]\]- Parameters
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import RosenbrockProblem, plot_2d_problem bounds = RosenbrockProblem.bounds # Contains traditional bounds plot_2d_problem(RosenbrockProblem(), xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)

-
bounds= (-2.048, 2.048)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued list phenome:
>>> phenome = [0.5, -0.2, 0.1] >>> RosenbrockProblem().evaluate(phenome) 22.3
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness
-
worse_than(first_fitness, second_fitness)¶ We minimize by default:
>>> s = NoisyQuarticProblem() >>> s.worse_than(100, 10) True
>>> s = NoisyQuarticProblem(maximize=True) >>> s.worse_than(100, 10) False
-
class
leap_ec.real_rep.problems.ScaledProblem(problem, new_bounds, maximize=None)¶ Scale the search space of a fitness function up or down.
-
evaluate(phenome)¶ Decode and evaluate the given individual based on its genome.
Practitioners must over-ride this member function.
Note that by default the individual comparison operators assume a maximization problem; if this is a minimization problem, then just negate the value when returning the fitness.
- Parameters
phenome –
- Returns
fitness
-
-
class
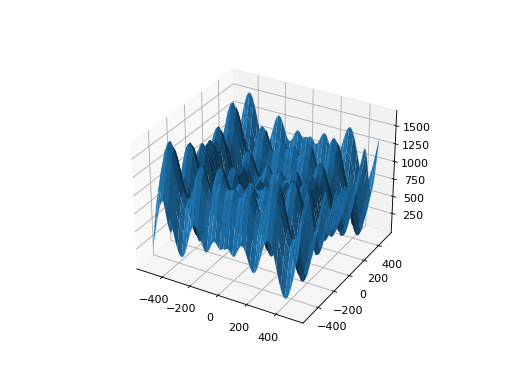
leap_ec.real_rep.problems.SchwefelProblem(alpha=418.982887, maximize=False)¶ Schwefel’s function is another traditional multimodal test function whose local optima are distributed in a slightly irregular way, and whose global optimum is out at the edge of the search space (with no gently sloping macrostructure to guide the algorithm toward it).
Compare this to the
RastriginProblemfunction, whose global optimum lies at the center of a quadratic bowl with a regular grid of local optima.\[f(\mathbf{x}) = \sum_{i=1}^d\left(-x_i \cdot\sin\left(\sqrt{\|x_i\|} \right)\right) + \alpha \cdot d\]- Parameters
alpha (float) – fitness offset (the default value ensures that the global optimum has zero fitness)
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import SchwefelProblem, plot_2d_problem bounds = SchwefelProblem.bounds # Contains traditional bounds plot_2d_problem(SchwefelProblem(), xlim=bounds, ylim=bounds, granularity=10)
(Source code, png, hires.png, pdf)
-
bounds= (-512, 512)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
class
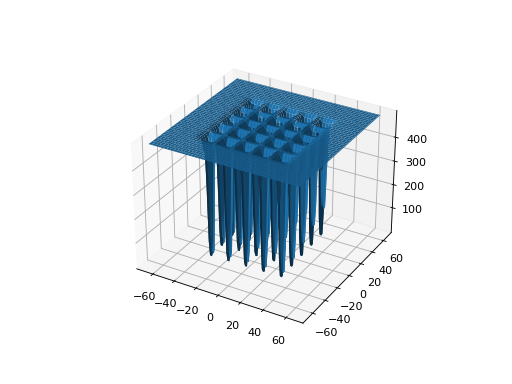
leap_ec.real_rep.problems.ShekelProblem(k=500, c=array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25]), maximize=False)¶ The classic ‘Shekel’s foxholes’ function.
\[f(\mathbf{x}) = \frac{1}{\frac{1}{K} + \sum_{j=1}^{25} \frac{1}{f_j(\mathbf{x})}}\]where
\[f_j(\mathbf{x}) = c_j + \sum_{i=1}^2 (x_i - a_{ij})^6\]and the points \(\left\{ (a_{1j}, a_{2j})\right\}_{j=1}^{25}\) define the functions various optima, and are given by the following hardcoded matrix:
\[\begin{split}\left[a_{ij}\right] = \left[ \begin{array}{lllllllllll} -32 & -16 & 0 & 16 & 32 & -32 & -16 & \cdots & 0 & 16 & 32 \\ -32 & -32 & -32 & -32 & -32 & -16 & -16 & \cdots & 32 & 32 & 32 \end{array} \right].\end{split}\]- Parameters
k (int) – the value of \(K\) in the fitness function.
c ([int]) – list of values for the function’s \(c_j\) parameters. Each c[j] approximately corresponds to the depth of the jth foxhole.
maximize (bool) – the function is maximized if True, else minimized.
maximize – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import ShekelProblem, plot_2d_problem bounds = ShekelProblem.bounds # Contains traditional bounds plot_2d_problem(ShekelProblem(), xlim=bounds, ylim=bounds, granularity=0.9)
(Source code, png, hires.png, pdf)
-
bounds= (-65.536, 65.536)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued list phenome (the output varies, since the function has noise).
- Parameters
phenome – real-valued to be evaluated
- Returns
its fitness
-
points= array([[-32, -16, 0, 16, 32, -32, -16, 0, 16, 32, -32, -16, 0, 16, 32, -32, -16, 0, 16, 32, -32, -16, 0, 16, 32], [-32, -32, -32, -32, -32, -16, -16, -16, -16, -16, 0, 0, 0, 0, 0, 16, 16, 16, 16, 16, 32, 32, 32, 32, 32]])¶
-
worse_than(first_fitness, second_fitness)¶ We minimize by default:
>>> s = ShekelProblem() >>> s.worse_than(100, 10) True
>>> s = ShekelProblem(maximize=True) >>> s.worse_than(100, 10) False
-
class
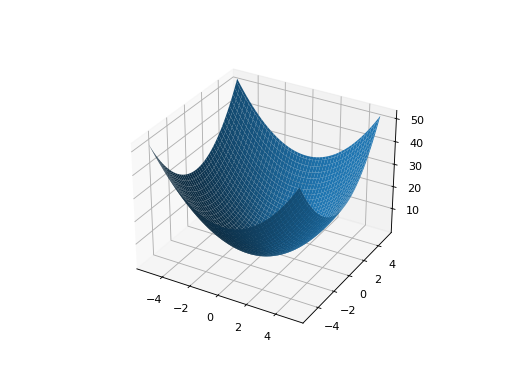
leap_ec.real_rep.problems.SpheroidProblem(maximize=False)¶ Classic paraboloid function, known as the “sphere” or “spheroid” problem, because its equal-fitness contours form (hyper)spheres in n > 2.
\[f(\vec{x}) = \sum_{i}^n x_i^2\]- Parameters
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import SpheroidProblem, plot_2d_problem bounds = SpheroidProblem.bounds # Contains traditional bounds plot_2d_problem(SpheroidProblem(), xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)
-
bounds= (-5.12, 5.12)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued list phenome:
>>> phenome = [0.5, 0.8, 1.5] >>> SpheroidProblem().evaluate(phenome) 3.14
- Parameters
phenome – real-valued vector to be evaluated
- Returns
it’s fitness, sum(phenome**2)
-
worse_than(first_fitness, second_fitness)¶ We minimize by default:
>>> s = NoisyQuarticProblem() >>> s.worse_than(100, 10) True
>>> s = NoisyQuarticProblem(maximize=True) >>> s.worse_than(100, 10) False
-
class
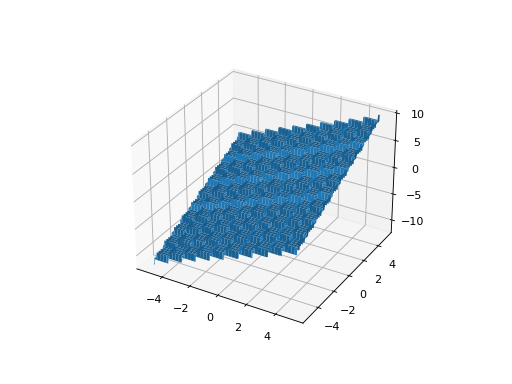
leap_ec.real_rep.problems.StepProblem(maximize=True)¶ The classic ‘step’ function—a function with a linear global structure, but with stair-like plateaus at the local level.
\[f(\mathbf{x}) = \sum_{i=1}^{n} \lfloor x_i \rfloor\]where \(\lfloor x \rfloor\) denotes the floor function.
- Parameters
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import StepProblem, plot_2d_problem bounds = StepProblem.bounds # Contains traditional bounds plot_2d_problem(StepProblem(), xlim=bounds, ylim=bounds, granularity=0.025)
(Source code, png, hires.png, pdf)
-
bounds= (-5.12, 5.12)¶
-
evaluate(phenome)¶ Computes the function value from a real-valued list phenome:
>>> phenome = [3.5, -3.8, 5.0] >>> StepProblem().evaluate(phenome) 4.0
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness
-
worse_than(first_fitness, second_fitness)¶ We maximize by default:
>>> s = StepProblem() >>> s.worse_than(100, 10) False
>>> s = StepProblem(maximize=False) >>> s.worse_than(100, 10) True
-
class
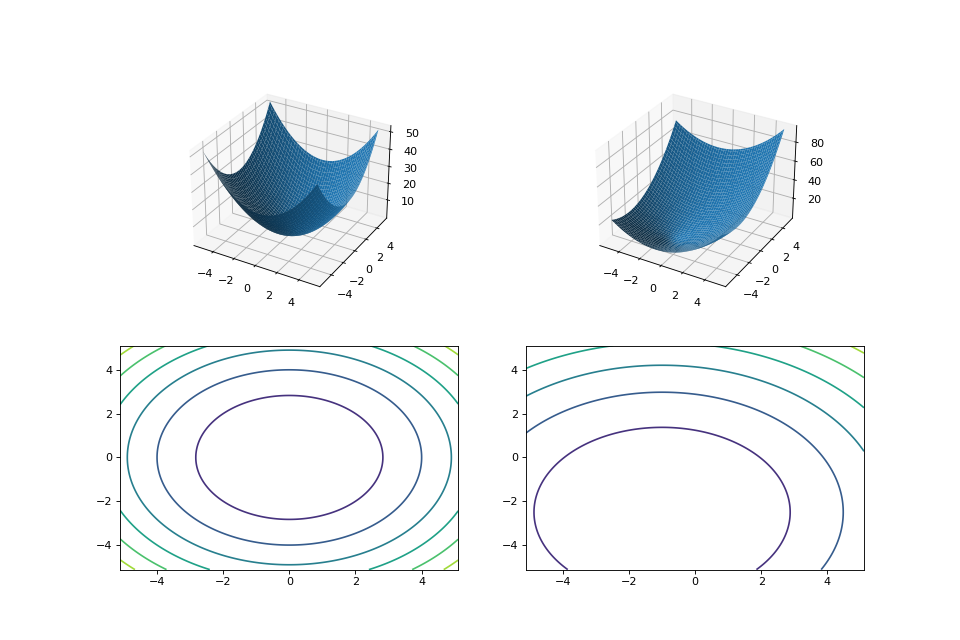
leap_ec.real_rep.problems.TranslatedProblem(problem, offset, maximize=None)¶ Takes an existing fitness function and translates it by applying a fixed offset vector.
For example,
from matplotlib import pyplot as plt from leap_ec.real_rep.problems import SpheroidProblem, TranslatedProblem, plot_2d_problem original_problem = SpheroidProblem() offset = [-1.0, -2.5] translated_problem = TranslatedProblem(original_problem, offset) fig = plt.figure(figsize=(12, 8)) plt.subplot(221, projection='3d') bounds = SpheroidProblem.bounds # Contains traditional bounds plot_2d_problem(original_problem, xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(222, projection='3d') plot_2d_problem(translated_problem, xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(223) plot_2d_problem(original_problem, kind='contour', xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025) plt.subplot(224) plot_2d_problem(translated_problem, kind='contour', xlim=bounds, ylim=bounds, ax=plt.gca(), granularity=0.025)
(Source code, png, hires.png, pdf)
-
evaluate(phenome)¶ Evaluate the fitness of a point after translating the fitness function.
Translation can be used in higher than two dimensions:
>>> offset = [-1.0, -1.0, 1.0, 1.0, -5.0] >>> t_sphere = TranslatedProblem(SpheroidProblem(), offset) >>> genome = [0.5, 2.0, 3.0, 8.5, -0.6] >>> t_sphere.evaluate(genome) 90.86
-
classmethod
random(problem, offset_bounds, dimensions, maximize=None)¶ Apply a random real-valued translation to a fitness function, sampled uniformly between min_offset and max_offset in every dimension.
from leap_ec.real_rep.problems import RastriginProblem, plot_2d_problem original_problem = RastriginProblem() bounds = RastriginProblem.bounds # Contains traditional bounds translated_problem = TranslatedProblem.random(original_problem, bounds, 2) plot_2d_problem(translated_problem, kind='contour', xlim=bounds, ylim=bounds)
-
-
class
leap_ec.real_rep.problems.WeierstrassProblem(kmax=20, a=0.5, b=3, maximize=False)¶ The Weierstrass function is famous for being the first discovered example of a function that is continuous, but not differentiable. Built by adding the terms of a Fourier series, it has a jagged, self-similar structure:
\[f(\mathbf{x}) = \sum_{i=1}^d \left[ \sum_{k=0}^{kmax} a^k \cos\left( 2\pi b^k(x_i + 0.5)\right) - n \sum_{k=0}^{kmax} a^k \cos(\pi b^k) \right]\]When used in optimization benchmarks, it’s typical to carry out the Fourier sum to kmax=20 terms.
- Parameters
kmax (int) – number of terms to carry the Fourier sum out to
a (float) – amplitude parameter of the cosine terms
b (float) – wavenumber (frequency) parameter of the cosine terms
maximize (bool) – the function is maximized if True, else minimized.
from leap_ec.real_rep.problems import WeierstrassProblem, plot_2d_problem bounds = WeierstrassProblem.bounds # Contains traditional bounds plot_2d_problem(WeierstrassProblem(), xlim=bounds, ylim=bounds, granularity=0.01)
(Source code, png, hires.png, pdf)

-
bounds= [-0.5, 0.5]¶
-
evaluate(phenome)¶ Computes the function value from a real-valued phenome.
- Parameters
phenome – real-valued vector to be evaluated
- Returns
its fitness.
-
leap_ec.real_rep.problems.plot_2d_contour(fun, xlim, ylim, granularity, ax=None)¶ Convenience method for plotting contours for a function that accepts 2-D real-valued inputs and produces a 1-D scalar output.
- Parameters
fun (function) – The function to plot.
xlim ((float, float)) – Bounds of the horizontal axes.
ylim ((float, float)) – Bounds of the vertical axis.
ax (Axes) – Matplotlib axes to plot to (if None, a new figure will be created).
granularity (float) – Spacing of the grid to sample points along.
The difference between this and
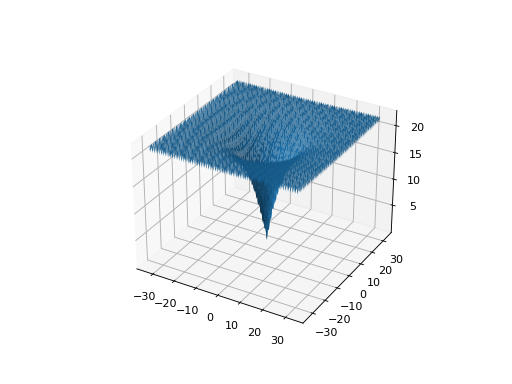
plot_2d_problem()is that this takes a raw function (instead of aProblemobject).import numpy as np from scipy import linalg from leap_ec.real_rep.problems import plot_2d_contour def sinc_hd(phenome): r = linalg.norm(phenome) return np.sin(r)/r plot_2d_contour(sinc_hd, xlim=(-10, 10), ylim=(-10, 10), granularity=0.2)
(Source code, png, hires.png, pdf)

-
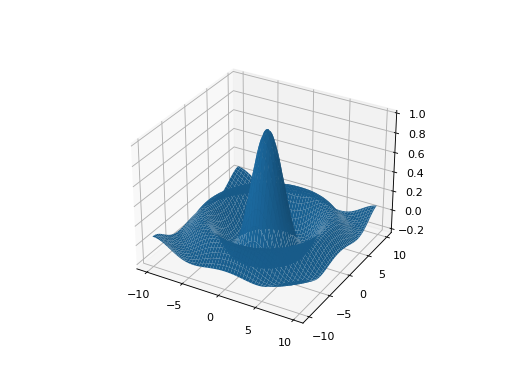
leap_ec.real_rep.problems.plot_2d_function(fun, xlim, ylim, granularity=0.1, ax=None)¶ Convenience method for plotting a function that accepts 2-D real-valued imputs and produces a 1-D scalar output.
- Parameters
fun (function) – The function to plot.
xlim ((float, float)) – Bounds of the horizontal axes.
ylim ((float, float)) – Bounds of the vertical axis.
ax (Axes) – Matplotlib axes to plot to (if None, a new figure will be created).
granularity (float) – Spacing of the grid to sample points along.
The difference between this and
plot_2d_problem()is that this takes a raw function (instead of aProblemobject).import numpy as np from scipy import linalg from leap_ec.real_rep.problems import plot_2d_function def sinc_hd(phenome): r = linalg.norm(phenome) return np.sin(r)/r plot_2d_function(sinc_hd, xlim=(-10, 10), ylim=(-10, 10), granularity=0.2)
(Source code, png, hires.png, pdf)
-
leap_ec.real_rep.problems.plot_2d_problem(problem, xlim, ylim, kind='surface', ax=None, granularity=None)¶ Convenience function for plotting a
Problemthat accepts 2-D real-valued phenomes and produces a 1-D scalar fitness output.- Parameters
fun (Problem) – The
Problemto plot.xlim ((float, float)) – Bounds of the horizontal axes.
ylim ((float, float)) – Bounds of the vertical axis.
kind (str) – The kind of plot to create: ‘surface’ or ‘contour’
ax (Axes) – Matplotlib axes to plot to (if None, a new figure will be created).
granularity (float) – Spacing of the grid to sample points along. If none is given, then the granularity will default to 1/50th of the range of the function’s bounds attribute.
The difference between this and
plot_2d_function()is that this takes aProblemobject (instead of a raw function).If no axes are specified, a new figure is created for the plot:
from leap_ec.real_rep.problems import CosineFamilyProblem, plot_2d_problem problem = CosineFamilyProblem(alpha=1.0, global_optima_counts=[2, 2], local_optima_counts=[2, 2]) plot_2d_problem(problem, xlim=(0, 1), ylim=(0, 1), granularity=0.025);
(Source code, png, hires.png, pdf)

You can also specify axes explicitly (ex. by using ax=plt.gca(). When plotting surfaces, you must configure your axes to use projection=’3d’. Contour plots don’t need 3D axes:
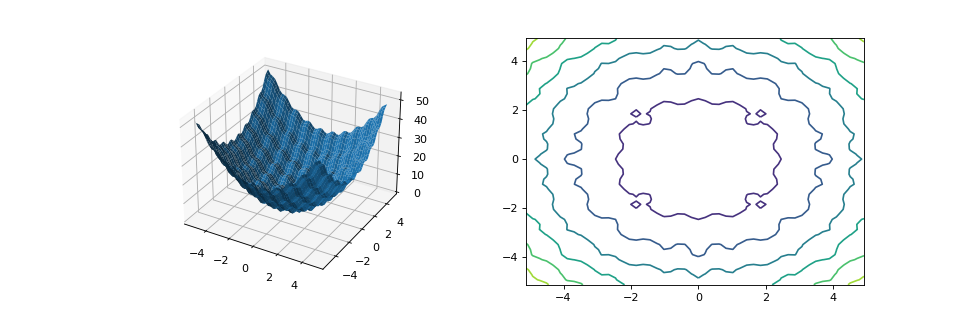
from matplotlib import pyplot as plt from leap_ec.real_rep.problems import RastriginProblem, plot_2d_problem fig = plt.figure(figsize=(12, 4)) bounds=RastriginProblem.bounds # Contains default bounds plt.subplot(121, projection='3d') plot_2d_problem(RastriginProblem(), ax=plt.gca(), xlim=bounds, ylim=bounds) plt.subplot(122) plot_2d_problem(RastriginProblem(), ax=plt.gca(), kind='contour', xlim=bounds, ylim=bounds)
(Source code, png, hires.png, pdf)